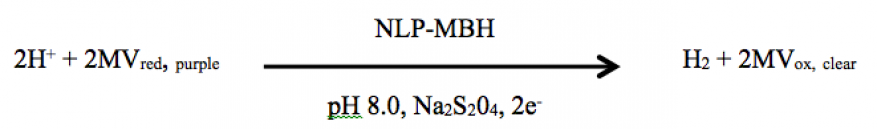Paul Hoeprich (16-FS-020)
Abstract
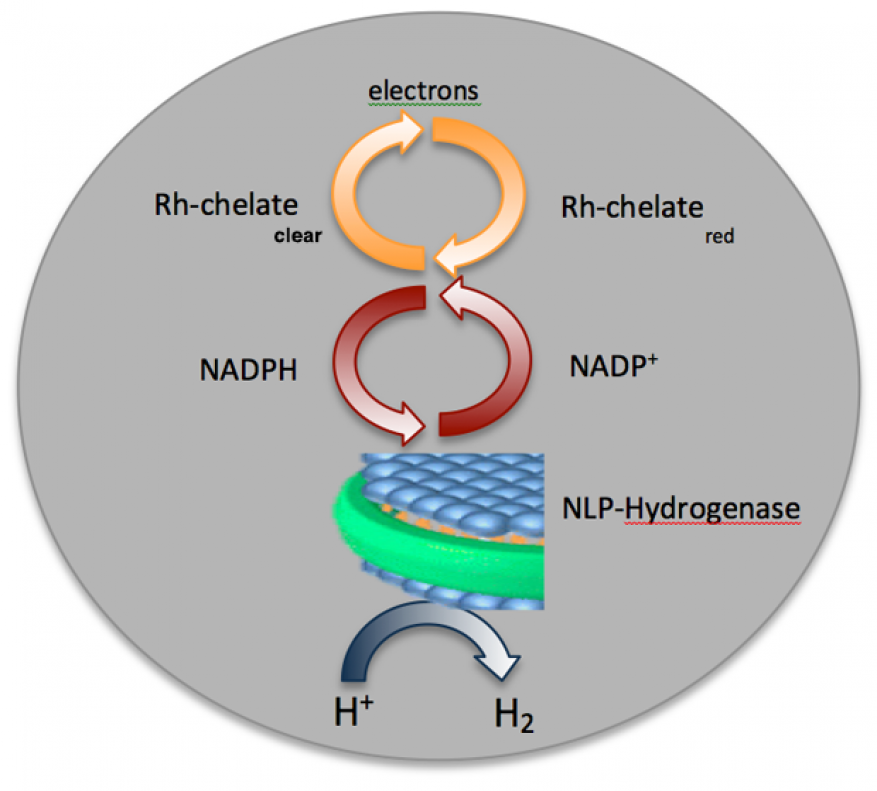
With the increasing demand for hydrogen transportation fuels, there is an urgent need to develop hydrogen gas production technology that is environmentally friendly, sustainable, and meets the ultimate promise of a clean and economical fuel. This approach stands in stark contrast to the dominant refinery-based steam-methane reforming technology that requires high temperature and pressure to convert methane into molecular hydrogen gas (H2). As hydrogen is produced naturally by various microorganisms, an attractive biological alternative to current production methods would be to use microbial hydrogenase enzymes to convert protons (H+) in aqueous solution to hydrogen gas under ambient conditions. In our project, we isolate two different bacterial membrane-associated hydrogenases and stabilize the enzymes in nanolipoprotein particle-based nanoconstructs. These nanolipoprotein particle–membrane-associated hydrogenase preparations are then tested in vitro for the production of H2 gas. If successful in the first instance, nanolipoprotein particle–membrane-associated hydrogenases would be incorporated into a flow-cell system that promotes the transfer of electrons from a small battery through a redox pathway to reduce H+ to H2 gas, using a unique rhodium catalyst to recycle the hydrogenase cofactor (nicotinamide adenine dinucleotide phosphate, or NADP+), as indicated in Figure 1.
Background and Research Objectives
In previous studies, we found that H2 is generated in vitro by partially purified nanolipoprotein particle–membrane-associated hydrogenase constructs. This was carried out in the presence of methyl viologen, a non-physiological electron carrier, which changes color from purple to clear when electrons are used by the membrane-associated hydrogenase enzyme to reduce 2H+ to H2.1 Because this membrane-associated hydrogenase contains an Fe-Fe reaction core, it is very sensitive to oxygen, necessitating cumbersome anaerobic work. In the present case, our objective was to repeat these experiments using Ni-Fe type membrane-associated hydrogenase enzymes, which are not sensitive to oxygen, from bacteria Cupriavidus metallidurans and Ralstonia spp.2,3 The reaction is:
Following the anticipated color change, hydrogen is measured by gas chromatography.
Our objectives for the six-month feasibility study were as follows:
- Prepare membrane fractions from two microbial species containing the Ni-Fe membrane-associated hydrogenase enzyme;
- Assemble, purify, and characterize nanolipoprotein particle–membrane-associated hydrogenase nanoconstructs;
- Measure H2 production by colorimetric assays (methyl viologen oxidation) and gas chromatography analysis.
Approach & Accomplishments
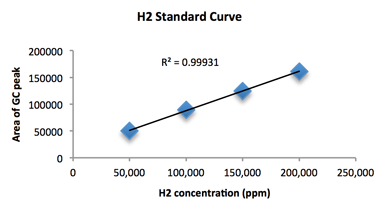
In FY16, we (1) re-furbished an older gas chromatography unit, which took about three months because it was an instrument that had been used specifically for H2 quantification in an earlier hydrogen production project, but had been idle for several years; (2) generated a H2 standard curve, against which our experimental samples could be compared and H2 production calculated (Figure 2); (3) prepared bacterial cell membranes (that should contain membrane-associated hydrogenases) following procedures established by previous investigators in the hydrogenase field4, which is to say, cells of both types of bacteria were grown to a specified density and then lysed using a French pressure cell; (4) collected membrane fractions by centrifugation and stored them in small aliquots at -80 °C; (5) generated H2 by simply mixing crude microbial membranes with dithionite (Na2S204), an electron donor, and the methyl viologen electron carrier; (6) monitored the redox reaction with methyl viologen, i.e., when the purple solution cleared, electrons were being transferred—presumably to H+ and enzymatic generation of H2; (7) made observations. Specifically, although we observed color loss in reactions with membrane preparations from both bacteria, and some gas chromatography measurements looked promising, we were unable to consistently detect H2 by gas chromatography. In addition, when cell membrane fragments were formulated into nanolipoprotein particles, we did not detect H2 evolution by gas chromatography.
Impact on Mission
At this point, impact is minimal. We were unable to demonstrate robust biological H2 production under ambient conditions, so the impact to Livermore’s hydrogen energy-related program will not be augmented or impacted.
Conclusion
Overall, this set of feasibility experiments failed to produce measureable H2. As to next steps, we believe the following:
- H2 might have been produced in amounts that caused feedback inhibition of membrane-associated hydrogenases, in either crude membranes or nanolipoprotein particle–membrane-associated hydrogenase nanoconstructs. Therefore, seeking removal of H2 immediately from reaction mixture might be important, e.g., binding to aerogel storage material or continuous flow to remove H2 from the reaction.
- Conduct experiments in a Coy chamber in an argon atmosphere—although we sought to circumvent the problem of O2 sensitivity by using a tolerant membrane-associated hydrogenase, the lack of O2 might enable H2 to persist and gas chromatography measurements become possible.
- Ni/Fe membrane-associated hydrogenases might not be as active as Fe/Fe hydrogenases towards proton reduction, but was appropriate for the experimental work performed. It would be interesting to try other potentially more active membrane-associated hydrogenase enzymes from other microbes.
References
- Baker, S. E., et al., “Hydrogen production by a hyperthermophilic membrane-bound hydrogenase in water-soluble nanolipoprotein particles,” JACS Comm. 131, 7508 (2009). LLNL-JRNL-408807. http://dx.doi.org/10.1021/ja809251f
- Cupriavidus metallidurans (ATCC 43123D-5).
- Goldet, G., et al., “Hydrogen production under aerobic conditions by membrane-bound hydrogenases from Ralstonia species.” Am. Chem. Soc. 130, 11106 (2008). http://dx.doi.org/10.1021/ja8027668
- Adams, M. W. W., et al. “Hydrogenase.” Biochem. Biophys. Acta 594(2–3), 105 (1981). http://dx.doi.org/10.1016/0304-4173(80)90007-5