Monica Borucki (13-ERD-020)
Abstract
A large proportion of infectious diseases go undiagnosed, which severely hampers prompt and appropriate treatment and containment. Recent advances in DNA sequencing-based characterization have advanced this field; however these techniques are limited by overwhelming backgrounds of host genome and commensal microbial flora. This is particularly problematic for RNA viruses, the most common cause of emergent disease, due to their small genome size. A possible solution could leverage the inherent RNA-binding capacity of the host innate immune response. Viral nucleic acid in particular is bound by numerous pathogen recognition receptors, which could provide a platform for selective pathogen enrichment. As a model system for this approach, BW5147 T lymphocyte cells were infected with Sindbis virus. Following incubation, the receptors TLR-3, MDA5, and RIG-I were extracted from cell lysates via immunoprecipitation. RNA fragments that were bound to pattern recognition receptors (PRR-bound) were extracted from the precipitated fraction, reverse transcribed, and subjected to whole-genome next-generation sequencing. Data were processed using the Livermore Metagenomics Analysis Toolkit. Results demonstrated a dramatic reduction in the relative quantity of host nucleic acid, and enrichment of Sindbis virus RNA in the precipitated fractions. Extraction using all three pathogen recognition receptors above yielded data for each region of the Sindbis virus genome. It was further observed that coupling with whole-genome amplification significantly improved coverage, which could facilitate genome assembly. Similarly, anti-IgM and anti-J chain antibodies were used in an immunoprecipitation assay to retrieve virus bound by immunoglobin (IgM) in lung lavage from mice infected with Sendai virus. However, although these results demonstrated successful enrichment of viral RNA through inherent innate immune mechanisms, the assay was periodically plagued by non-specific binding of the negative-control antibodies. Optimization of assay parameters was not able to overcome this problem rendering the assay results unreliable.
Background and Research Objectives
Novel pathogens may circulate in a population for years prior to detection. In fact, recognition of past outbreaks of emergent pathogens has been delayed due to lack of biosurveillance. Efficient biosurveillance techniques need to be developed to avoid unexpected epidemics and rapidly detect novel diseases such as HIV, New World hantaviruses, avian influenza, Severe Acute Respiratory Syndrome virus, and Middle East Respiratory Syndrome virus. It is estimated that 69% of upper respiratory diseases go undiagnosed,1 noteworthy because respiratory diseases that are efficiently transmitted have the greatest potential to result in pandemic disease.
Currently, the process of novel pathogen detection and characterization is constrained due to the difficulties of isolating and characterizing unique microbes for which there are no reagents available and which are not readily cultivated using conventional techniques. Recent advances in genetic characterization of microbes include microarrays, metagenomics, and next-generation sequencing platforms. However, these techniques rely on detecting the microbe genome, which exists in much smaller amounts compared to the host genome and commensal microbial flora. This is particularly problematic for novel viruses, which have small genomes, such as RNA viruses, the most common cause of emergent diseases.2,3
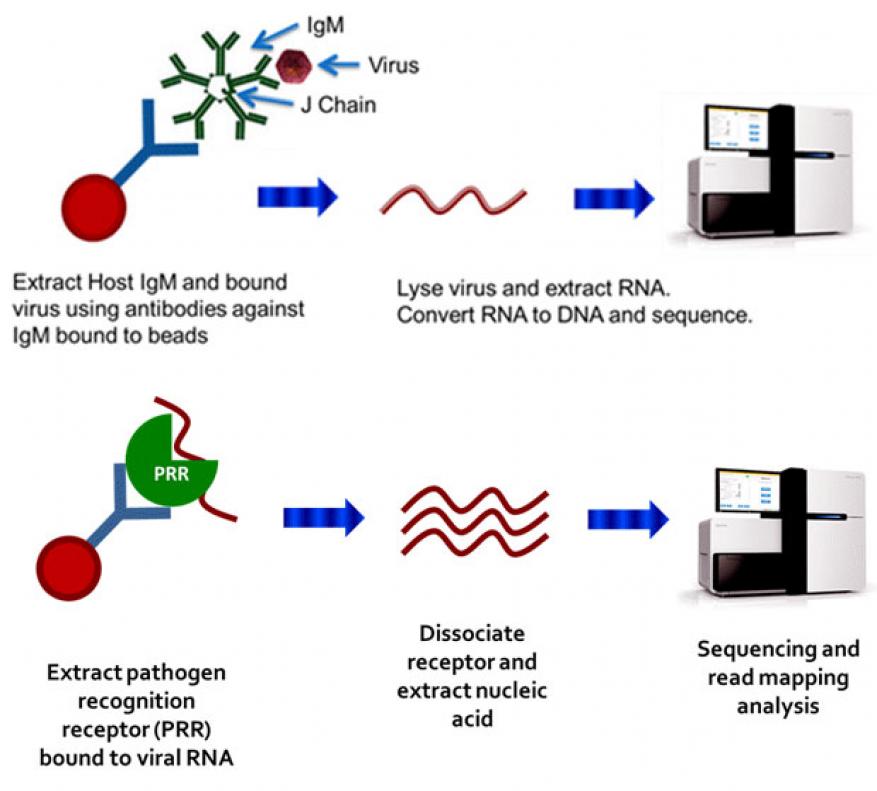
The host innate immune response has evolved mechanisms for identifying non-self entities and sensing pathogens. An invading microbe triggers nonspecific humoral and cellular immune responses, with the antigen being bound by circulating IgM and pathogen-recognition receptors such as Toll-like receptors (TLRs).4,5 Viral nucleic acid is bound in particular by RIG-I, MDA5, TLR3, and TLR9, which play major roles in microbial recognition and interferon induction. We hypothesized that immunoprecipitation using anti-pathogen-recognition receptor antibodies or anti-IgM antibodies can be used to enrich for viral RNA bound by pathogen-recognition receptors or viral particles (virions) bound by IgM, thus allowing viral nucleic acid to be concentrated away from contaminating host nucleic acid and permitting subsequent metagenomic characterization of the pathogen genome (Figure 1, top and bottom, respectively).
Scientific Approach and Accomplishments
The goal of this project was to develop relatively simple techniques that separate the nucleic acid of novel pathogens from the overwhelming genetic material of host and background flora. This was accomplished by leveraging the body’s earliest and pathogen-agnostic immune response to infection. Methods were developed to rapidly isolate pathogen genetic material or intact pathogens that are captured by the host pathogen-recognition receptors and IgM response, respectively. Initially Western blots were used to confirm that the antibodies used in the assay to capture the target proteins (pathogen-recognition receptors, IgM) were indeed binding the antigen. Subsequently, IP assays were developed using magnetic beads coated with anti-pathogen-recognition receptor antibodies or anti-IgM antibodies to capture the viral RNA bound by pathogen-recognition receptors or virions bound by IgM. Both the bound fraction and the unbound fractions were tested for the presence of host 18S ribosomal RNA and viral genomic RNA to determine if the host RNA was selectively being removed. Because the IgM assay captured the virion, and thus the entire viral genome intact, quantitative reverse transcription polymerase chain reactions could be used to determine the ratio of host RNA to the viral RNA, and thus assess the assay performance. However, pathogen-recognition receptors bind only fragments of the viral genome ranging in length from about 200 nt to 2,000 nt and the bound fragments can originate from any region of the genome,6,7 thus it was not possible to accurately gauge assay performance of the pathogen-recognition receptors assay prior to metagenomic analysis. To address this, a highly multiplexed polymerase chain reaction assay with primers scanning the entire Sindbis genome was designed by Livermore bioinformaticist Shea Gardner, with polymerase chain reaction primers targeting every 200 nt region of the genome.8This enabled us to assess changes in yield of polymerase chain reaction products and select the most productive protocol for use (Figure 2). Once the assay output was optimized, samples were prepared for metagenomic analysis to definitively determine assay performance. Initial results were promising; however yields of retained RNA were low and an additional step was added to nonspecifically amplify the nucleic acid via whole genome amplification using the phi29-based Repli-g polymerase (Qiagen). The viral and host genomes captured using these techniques were characterized using metagenomics and computational analysis to identify the organisms present in the sample. Once these methods were tested and optimized on control samples, these were applied to human and veterinary samples.
Results
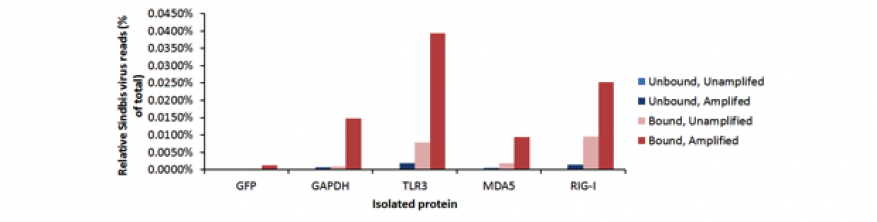
As a model system for the pathogen-recognition receptor approach, BW5147 T cells were infected with Sindbis virus. This cell line was selected because it exhibits robust expression of each of the pathogen-recognition receptors of interest, and is amenable to Sindbis virus infection. Following incubation, receptors TLR-3, MDA5, and RIG-I were extracted from cell lysates via immunoprecipitation. Antibodies against green fluorescent protein or host metabolic protein glyceraldehyde-3-phosphate dehydrogenase (GAPDH) were employed as a control, as these proteins should not exhibit any binding to viral RNA. Western blots were used to validate the pathogen-recognition receptors immunoprecipitation data, and highly multiplexed polymerase chain reactions were used to show the presence of viral RNA associated with the pathogen-recognition receptors (Figure 2). The DNA bands obtained from “bound” samples represents nucleic acid co-extracted with the pathogen-recognition receptors of interest, while the “unbound” samples contain all nucleic acid not associated with pathogen-recognition receptors. A smaller fraction of the total RNA is present in “bound” fractions (as represented by fainter bands in Figure 2), as these receptors bind only specific nucleic acid motifs particular to pathogens. PRR-bound RNA fragments were extracted from the bound and unbound fractions and subjected to next-generation Illumina sequencing. Data were processed using the Livermore Metagenomics Analysis Toolkit.9 Results demonstrated a dramatic reduction in the relative quantity of host nucleic acid, and enrichment of viral RNA in the precipitated fractions (Figure 3). This enrichment was observably pronounced over results obtained with the green fluorescent protein control.
Despite the observed enrichment, total read counts for each sample were relatively low across samples. Non-selective whole genome amplification was therefore applied in an attempt to increase the dynamic range of detection. Coupling the immunoprecipitation process with whole genome amplification further improved the observed enrichment between bound and unbound fractions.
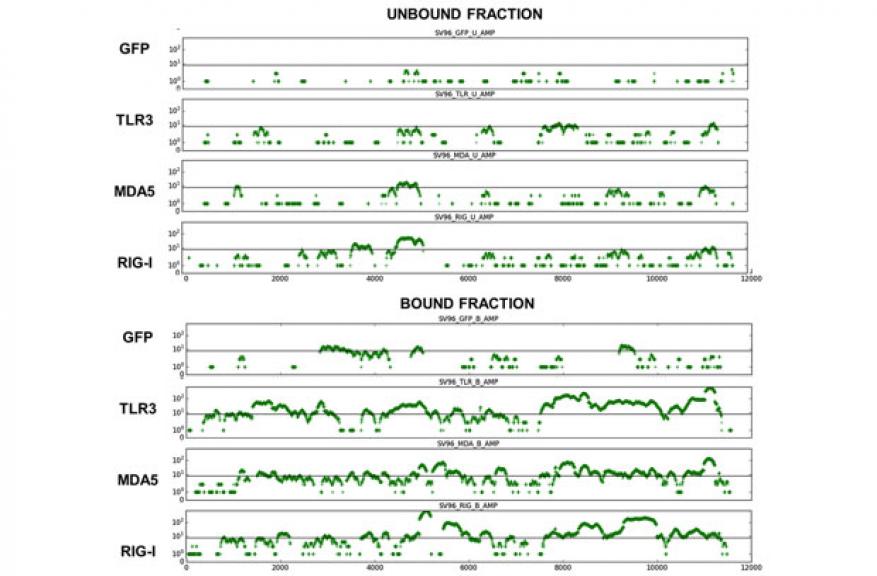
Extracted sequence reads were mapped to different portions of the viral genome to determine the coverage of observed reads. Extraction yielded data for each region of the viral genome, though certain regions of the genome exhibited higher coverage, indicating preferred binding points, which may be reflective of the primary and secondary structure of RNA from these regions (Figure 4). The improvement in coverage when coupled with whole genome amplification could facilitate genome assembly toward identification of novel viruses.
Although viral RNA enrichment increased the number of viral reads generated, non-specific binding was persistently a problem. Initially, antibodies to host metabolic protein GAPDH were used as a control. However, the GAPDH antibodies did bind to viral nucleic acid (Figure 3) and a review of literature revealed that other studies had similar findings.10 Two other negative control antibodies were tried including one, green fluorescent protein, which has worked in a similar protocol.11 Numerous optimization efforts were applied, including attempts to block non-specific binding sites with a multitude of protein candidates, altering incubation time, adjusting assay temperature, and modifying stringency of the incubation buffer. Despite protocol optimization steps, however, the method was plagued by non-specific binding of the negative control antibodies and analysis of clinical samples also gave inconsistent results, both between biological and technical replicates. It is likely that the proportion of RNA bound to pathogen-recognition receptors within the cell is a very small fraction of total viral RNA within the cell lysate. Thus the observed signal is challenging to identify above the general noise of non-specific RNA binding to control proteins or polymer matrix comprising the bead support system. This low signal-to-noise ratio was manifested in a high degree of variation in observed results. Engineering a higher infection titer, or higher pathogen-recognition-receptor-to-virion ratio may be required to achieve observably consistent results, which may not be possible within a clinical context.
Capture of virus bound to IgM from sera samples obtained from mice infected with virus was assayed using IgM-binding columns and beads. Amounts of host 18S RNA and viral RNA present before and after treatment were measured using quantitative reverse transcription polymerase chain reaction. Preliminary experiments determined that at least three logs of host nucleic acid were removed using the column and the bead assays. Nuclease treatment was also tested for reduction of host nucleic acid as a separate step after IgM capture but was found to adversely affect yield of viral nucleic acid.
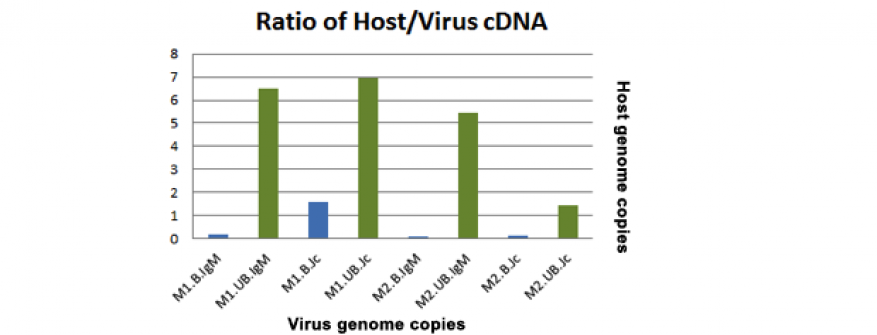
Although a significant amount of host RNA was removed, results of the quantitative reverse transcription polymerase chain reaction and subsequent metagenomic assays indicated that saturation of the antibody binding sites reduced recovery of viral RNA from sera due to the large amount of unbound IgM in sera. To increase the sensitivity of the IgM assay we sought to optimize the assay by using capture antibodies that favor binding of IgM antibodies that are bound to antigen (IgM shifts shape when bound to antigen), such as antibodies that bind the internal J chain of IgM. The J chain is a structural support for the IgM molecule, and access to this portion should be exposed to a greater degree when the molecule is bound to a ligand. We also changed the clinical sample tested from sera to mucosal secretions such as lung lavage or nasal samples in an effort to increase the ratio of bound to unbound IgM present in the sample. Immunoprecipitation assays using IgM and J chain were tested using lung lavage from mice infected with Sendai virus and yielded promising polymerase chain reaction data (Figure 5) but inconsistent metagenomic data (1 out of 2 experiments detected a high number of Sendai virus reads whereas the second did not). Nasal samples from camels infected with Middle East Respiratory Syndrome coronavirus were obtained from Dr. Richard Bowen at Colorado State University and assayed using anti-bovine IgM antibodies to capture the camel IgM (versus anti-mouse antibodies as used previously). These samples were an excellent test case, as they represented clinically oriented samples derived from a novel viral infection. Polymerase chain reaction data indicated that the assays were successful, but metagenomics data showed few Middle East Respiratory Syndrome reads, possibly because of the use of an anti-bovine IgM capture antibody, since antibodies specific to camel IgM were not available. As a control, the MERS-infected nasal samples were subject to Illumina sequencing, and the data indicated extreme genetic diversity of the Middle East Respiratory Syndrome genome present in the samples.
Impact on Mission
Although this assay format did not yield consistent enough results to be further developed, a number of new capabilities and collaborations resulted from the effort. In particular, scientific staff were trained in virology, cell culture, preparation of Illumina sequence libraries, and bioinformatics techniques, thus increasing the depth of Livermore’s capacity to work on future projects involving emerging viral pathogens. These capabilities were applied to the novel Middle East Respiratory Syndrome virus, and genomic data with high value to the scientific community were collected and published. Ability to process and work with this virus, as well as the corresponding publication record, represents value and sponsor interest for future work.
Conclusion
Immunoprecipitation assays were developed for multiple pathogen-recognition receptors and tested using BW 5147 cells infected with Sindbis virus. Metagenomic data showed the assay removed over 90% of host nucleic acid. By using multiple pathogen-recognition receptors in the assay, we were able to obtain sequence data for all regions of the viral genome. The low fraction of total viral RNA bound to PRRs in a cellular context, however, was demonstrated as likely below the limit of detection in a clinical context, rendering the assay to be of limited practical applicability in its current form. Collaborations were formed as a result of this work involving Colorado State University, which resulted in a publication, and the University of California, Davis, which led to joint submission of a grant proposal.
References
- Lodes, M. J., et al. “Identification of upper respiratory tract pathogens using electrochemical detection on an oligonucleotide microarray.” PLoS ONE 2, e924 (2007).
- Katze, M. G., et al., “Innate immune modulation by RNA viruses: Emerging insights from functional genomics.” Nat. Rev. Immunol. 8, 644 (2008).
- Nichol, S. T., Arikawa, J., and Kawaoka, Y. “Emerging viral diseases.” Proc. Nat. Acad. Sci. 97, 12411 (2000).
- Kawai, T., and Akira, S. “Toll-like receptors and their crosstalk with other innate receptors in infection and immunity.” Immunity 34, 637 (2011).
- Loo, Y.-M., et al. “Distinct RIG-I and MDA5 signaling by RNA viruses in innate immunity.” J. Virology 82, 335 (2008).
- Jensen, S., and A. R. Thomsen, “Sensing of RNA viruses: a review of innate immune receptors involved in recognizing RNA virus invasion.” J. Virology 86, 2900 (2012).
- Kato, H., et al., “Length-dependent recognition of double-stranded ribonucleic acids by retinoic acid–inducible gene-I and melanoma differentiation–associated gene 5.” J. Exp. Med. 205, 1601 (2008).
- Be, N. A., et al., “Microbial profiling of combat wound infection through detection microarray and next-generation sequencing.” J. Clin. Microbiol. 52, 2583 (2014).
- Ames, S. K., et al., “Scalable metagenomic taxonomy classification using a reference genome database.” Bioinformatics 29, 2253 (2013).
- Yi, M., D. E. Schultz, and S. M. Lemon, “Functional significance of the interaction of hepatitis A virus RNA with glyceraldehyde 3-phosphate dehydrogenase (GAPDH): Opposing effects of GAPDH and polypyrimidine tract binding protein on internal ribosome entry site function.” J. Virology 74, 6459 (2000).
- Baum, A., R. Sachidanandam, and A. García-Sastre, “Preference of RIG-I for short viral RNA molecules in infected cells revealed by next-generation sequencing.” Proc. Natl. Acad. Sci. Unit. States Am. 107(37), 16303 (2010).
Publications and Presentations
- Borucki, M., et al., Middle East respiratory syndrome coronavirus intra-host populations are characterized by numerous high frequency variants. American Society for Microbiology Biodefense and Emerging Diseases Research Mtg. Arlington, VA, Feb. 8–10, 2016. LLNL-ABS-678358.