Bradley Hart (16-SI-002)
Project Description
Traditional mainstays of forensic science that depend on qualitative expert opinion such as hair and fingerprint analysis are being challenged regularly as lacking a scientific and statistical basis. These challenges, leading to questions of admissibility of this type of evidence in current cases and validity of previous convictions, are potentially leading to a state of crisis in forensics. The shortcomings of these qualitative methods not only undermine the criminal justice system, but also increase resistance to their use in the broader national security community. The deoxyribonucleic acid (DNA) typing methodology stands out as an exception, but its use is limited when the sample has degraded or when there are multiple contributors. Protein can serve as a surrogate for DNA in these cases and provide an alternative quantifiable analysis strategy. Furthermore, genetically useful information can be obtained from genetically variant peptides extracted from protein samples. We plan to develop the scientific and statistical basis for forensic use of analyses based on genetically variant peptides and extend the concept into other forensic and intelligence contexts. We will focus on the tissue types and quantities commonly found at crime scenes and develop sample preparation and analytical methods to detect variation in the form of genetically variant peptides. The project has three broad objectives: (1) enable human identification from a single hair, (2) develop techniques for other forensically relevant tissue sources, and (3) exploit next-generation sequencing to obtain unique and specific peptides to track individuals and resolve complex mixtures. Each of these methods will incorporate automated processes that are compatible with microscopic- and nanometer-scale fluidic sample handling and analysis, and will require the development of novel bioinformatic and biostatistical methodologies. The methods we develop will also significantly impact biological archeology by providing biogeographic information or sex determination for samples that no longer contain usable DNA for these purposes. We will also develop strategies to track individuals, even against a complex background of many individuals, using genetically variant peptides related to ultrarare variants.
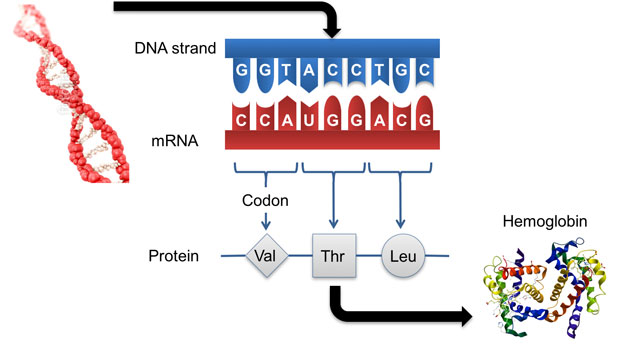
Forensic science is currently under intense scrutiny, particularly in the area of hair and fiber analysis, related to the reliability of subjective expert-witness testimony. Our research will directly address this issue by providing quantifiable analysis of hair, teeth, and bone evidence to replace subjective methods. By focusing on the tissue types commonly found at crime scenes, we will add protein typing as a scientific and statistically rigorous forensic tool. We expect to establish the technical foundation for using proteins as a source of information for human identification (see figure). Expected results include (1) integrating measures of identity based on genetically variant peptides and DNA; (2) developing a biogeographic prediction algorithm; (3) developing methods to identify, characterize, and validate genetically variant peptides expressed in bone and teeth, and evidenced in palmprints and fingerprints; (4) developing a methodology to identify sex-specific peptides; (5) identifying unique or rare genetically variant peptides in hair and palms; and (6) developing a workflow for identifying unique or rare DNA sequence variations (single-nucleotide polymorphisms) that occur commonly within a given population.
Mission Relevance
This goal of this effort is to provide a scientific and statistically validated methodology that will impact the fields of forensics, biological archeology, and proteomics. Our research directly addresses gaps in forensic science that have been nationally highlighted and acknowledged, and supports the strategic focus area of cyber security, space, and intelligence, particularly with respect to providing novel forensic methods with applications in counter-proliferation, counterterrorism, and domestic security, as well as expands the LLNL bioscience and bioengineering core competency.
FY16 Accomplishments and Results
In FY16 we (1) initiated collection and analysis of samples from a cohort of European American and East Asian human subjects; (2) performed analysis of both existing and new samples with the aims of optimizing sample processing and analysis methods to decrease sample size requirements as well as discovery and validation of new genetically variant peptides in hair protein; (3) achieved single-hair sample dissolution; (4) analyzed mass spectrometric data, resulting in 27 newly discovered genetically variant peptide candidates that were consistent with whole exome sequencing (using all the expressed genes in a genome, or exome) and detecting 28 already established and characterized genetically variant peptides; (5) identified multiple new potential genetically variant peptides in bone tissue; and (6) detected peptides unique to the sex-related isoforms (protein variants) of the amelogenin protein in tooth samples.
Publications and Presentations
- Anex, D., and G. Parker, Protein-based human identification. (2016). LLNL-POST-690338.
- Kiesow, C. W., et al., Detection of genetic information in the enamel proteome. (2016). LLNL-ABS-699919.
- Mason, K. E., et al., Sex-determination: A novel protein-based sex assignment technique using human tooth enamel and mass spectrometry. (2016). LLNL-ABS-699130.
- Parker, G. J., et al., "Demonstration of protein-based human identification using the hair shaft proteome." PLos One (2016). LLNL-JRNL-665656. http://dx.doi.org/10.1371/journal.pone.0160653