Elizabeth Wheeler (17-ERD-121)
Executive Summary
We are developing a three-dimensional experimental platform for use in characterizing tumor growth and drug response down to the sub-cellular level. This research will aid the development of medical countermeasures for rapid mitigation of evolving and unknown biological threats, and could lead to a new, integrated approach for rational cancer drug development.
Project Description
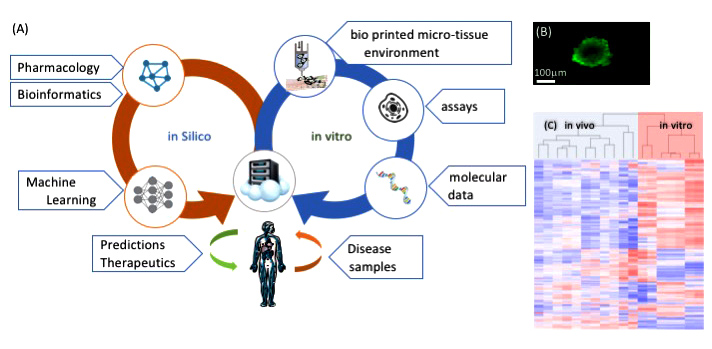
Defending against biological threats requires predictive disease models and therapeutic screens. The greatest challenge to advancing these technologies is the lack of adequate in vitro models that accurately reflect the complexity of the in vivo microenvironment. For instance, although much has been gleaned from studying two-dimensional human cancer cell lines, these studies offer little predictive value in understanding the complex relationship between tumor composition, microenvironment, and functional outcomes. One current, leading-edge approach combines state-of-the-art computational methods and high-performance computing to model cancer. However, the success of such an approach requires the availability of reliable, high-fidelity, experimental data to calibrate model parameters and validate model predictions. Addressing this challenge, we will develop an instrumented, human, ex vivo, three-dimensional tumor-model platform for unprecedented spatio-temporal control and measurement of multiple tissue, cellular, and sub-cellular components that drive tumor growth (see figure). We will recapitulate characterized tumors in three dimensions using biological printing (creating spatially controlled compartments that support biological cell function) with integrated, multi-scale sensing (the monitoring of multiple phenomena).
We intend to create a novel three-dimensional experimental platform for quantitative characterization of normal- and tumor-tissue response to interventions and demonstrate a new, integrated, experimental and computational approach for rational cancer drug development at scale. By blending innovative technologies with established science and technology expertise, we expect to acquire novel, large, and diverse data sets that will aid in modeling tumor growth and drug response, and assist in making personalized treatment choices. Working in close collaboration with academic, industrial, and government agency partners, we will characterize the tumor and its microenvironment including molecular, cellular, extracellular matrix, and stromal composition. This information will be used to recapitulate these tumors in three dimensions using biological printing to spatially control cellular compartments with integrated multi-scale sensing. Acquired data on response (extracellular and single cell) to a chemotherapeutic drug will be leveraged to instantiate an integrated experimental and computational approach to understanding tumor response. We expect this technology to advance the science of predictive disease modeling and the data to impact cancer biology by generating novel prognostic and therapeutic candidate biomarkers of tumor efficacy.
Mission Relevance
This research addresses the NNSA’s strategic goal of strengthening its science, technology, and engineering base by expanding and applying our science and technology capabilities to deal with national security challenges. Specifically, this project supports Livermore’s biosecurity mission in the area of medical countermeasures for rapid mitigation of evolving and unknown threats. The project is expected to have a broad impact, driving innovation across the Laboratory’s core competencies in bioscience, bioengineering, high-performance computing, computer-based simulation, and data science.
FY17 Accomplishments and Results
In the first four months of this project, which started late in the fiscal year, we focused on characterizing the cellular components of an in vivo tumor. In addition, we developed a compatible biological ink to successfully print viable cancer cells.