Felice Lightstone (16-FS-007)
Project Description
One of the most critical cellular processes that controls almost every aspect of life is the signaling that occurs from one side of a cell membrane to the other. However, lipid composition can alter the membrane fluidity, thickness, flexibility, and electrostatic properties. Additionally, different tissue types contain different compositions of lipids in their cell membranes. Thus, cell behavior, and therefore response to external stimulus, can vary in different parts of the body. Furthermore, recent studies postulate the existence of microscopic domains of certain lipid compositions within the cell membrane. It is therefore vitally important to understand how these lipid compositions, and thus membrane behavior, vary throughout the body. In addition, the toxicological outcome of taking a drug or toxicant is determined by its concentration in the body over time, and accurately predicting the distribution of a drug or toxicant in the human body is largely controlled by membrane permeability through the different tissues and organs. We plan to explore the feasibility of using coarse-grained and atomistic simulations to model human tissue membranes by changing the lipid composition. We will model lipid membranes to mirror the lipid composition of organs in the body and simulate the permeability of drugs and toxicants across those membranes. Simulations will be compared to experimental results to validate the minimal lipid compositions that accurately model the tissue types. This knowledge can have significant impact on the drug design and development process, which is estimated to cost about 2 billion dollars and take 15 years for a drug to reach market. During this process, 90% of drug candidates fail in clinical trials because of toxicological effects or being ineffective against their intended target disease.
We intend to explore the feasibility of using cutting-edge molecular dynamics simulations to computationally investigate how the variation in lipid composition present in different tissue types alters the properties of those membranes, and identify microscopic domains present within the membranes. We will simulate three different membrane patches, each with a tissue-specific lipid composition (pancreas, lung, and liver). Following equilibration of the lipid patches and formation of potential microscopic domains, representative subsections of each membrane will be converted into an atomistic-level model for more detailed refinement and analysis. Previous studies have shown aspects of permeation to be dependent on membrane composition. However, it is unclear if simply changing the lipid composition will be sufficient to model differences in tissue membranes and permeability into those tissues. The permeability of well-studied small molecules will be calculated through each membrane subsection using our established computational methodology. By combining the permeability properties of each membrane subsection, we will be able to constitute an overall permeability for the entire membrane patch, and thus generate a predicted permeability for that specific tissue. Simulated results will be compared to experimental results to validate the minimal lipid compositions that accurately model the tissue types. If successful, we expect to create accurate atomistic and coarse-grained lipid models, and validate their feasibility for future studies. In addition, the methodology to create these models and the protocols to validate the results will allow the development of models for additional human organs or for those of other species. The resulting models will also be used to simulate unknown drug candidates and toxicants to enable streamlined drug development efforts and toxicant identification. These simulations can improve current models of how the human body reacts to a drug to be more predictive of unknown compounds.
Mission Relevance
The creation of human tissue membrane models supports the Laboratory’s core competency in bioscience and bioengineering by determining the feasibility of computational testing and prediction to identify better drug candidates faster. We can also use these membrane models to simulate exposure to unknown chemicals to predict the human outcomes, relevant to chemical and biological security. Support of high-performance computing, simulation, and data science is realized through the development of new models and protocols that could be applied to other high-performance computing efforts. The anticipated size of the simulations is large and will require newly designed workflows for molecular dynamics simulations.
FY16 Accomplishments and Results
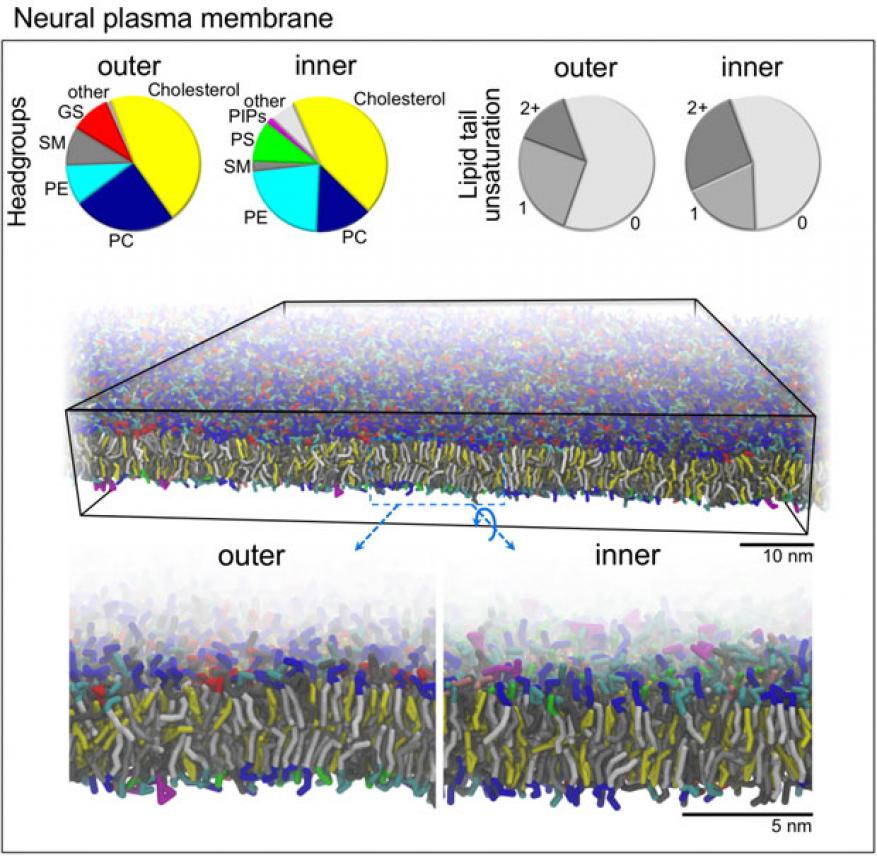
The feasibility of creating lipid compositions of different human organs has been quite challenging—a limited amount of data is available for human plasma membranes for specific organs. However, in FY16 we (1) created two models of a human brain-cell membrane composed of 16 and 60 different lipid types (see figure); (2) ran these models at a coarse-grained resolution using molecular dynamics simulations; (3) completed the coarse-grained simulations for human-brain cell membranes; and (4) converted the coarse-grained representation to an atomistic level for the 16-lipid model. The atomistic simulations will continue to run into the next fiscal year.